Hi!!
I am Deepankar, but I usually go by the shorter version: “Deep”. I have been working in the domain of Cancer research for over 11 years now. I have been involved in several projects, which were aimed at understanding molecular mechanisms of different human cancers and identifying opportunities for developing targeted therapeutics.
During the PhD, I developed a new high-throughput screen to discern activating mutations in kinases by constructing a cDNA library encompassing all possible point-mutations in the said kinase. So far, the screen has been used to study the EGFR family of RTKs (EGFR, HER2, HER3, and HER4) and identify activating mutations and assess their druggability with various TKIs. Using my screening workflow I am also investigating therapeutic resistance to ERBB-targeted inhibitors. During my spare time, I also developed a collection of webtools using R and shiny, some of which are publicly hosted at https://eleniuslabtools.utu.fi
In January 2023, I joined Genentech, Inc. as a Postdoctoral Research Fellow where I am currently studying WNT signaling in various molecular subtypes of Colorectal Cancer.
Here is my detailed CV in a PDF format.
- WNT signaling
- Colorectal Cancer
- Cancer Stemness
- Cell Differentiation
- Kinases & Kinase inhibitors
- Receptor Tyrosine Kinases
- Targeted therapeutics
- Therapeutic resistance
- Predictive biomarkers
- Somatic mutations in Cancer
PhD (Medical biochemistry & genetics), 2023
Institute of Biomedicine, University of Turku, Finland
MSc (Bioinformatics), 2014
Turku Centre for Biotechnology, University of Turku, Finland
B.Tech. (Biotechnology), 2012
Amity Institute of Biotechnology, Amity University, India
Experience
Conducting Functional Genetics research at the laboratory of Prof. Klaus Elenius. My responsibilities included:
- Planning and conducting Functional Genetics experiments
- Mammalian cell culture & dose-response analysis (Kinase inhibitors)
- Analysis of next-generation sequencing data
- Link to my PhD Thesis
Featured Publication
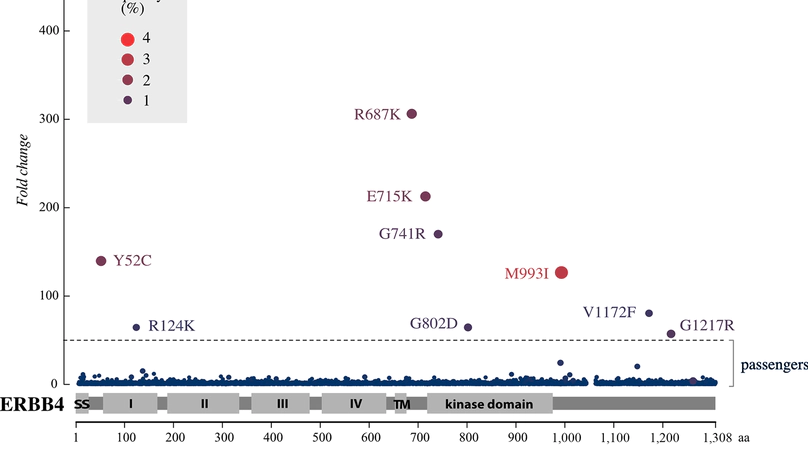
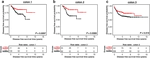
ERBB4 is a member of the ERBB family of oncogenes (with EGFR, HER2 & HER3). ERBB4 is frequently mutated in different cancers, but the functional impact of its somatic mutations remains unknown. Here, in an unbiased functional genetics screen, we analyzed the function of over 8,000 ERBB4 variants that were generated by random mutagenesis. The data indicate the presence of rare activating ERBB4 mutations in cancer, with potential to be targeted with clinically approved pan-ERBB inhibitors.
Recent Publications
Patents
Funding
Personal Grants
Conferences
Travel Grant
- Syöpäjärjestöt Travel grant
- TuDMM Travel grant
- LIFESPAN research program, Turku Travel grant
Travel Grant
- Turku Doctoral programme for Molecular Medicine
- LIFESPAN research program, Turku
Travel grant
- Turku Doctoral programme for Molecular Medicine
- Reagent & Experiments covered by EMBO.
- Accomodation (partial) sponsored by EMBO partners.
Hobbies
I ❤️ Primes!
Swim & Snorkel
Shell, R, Python
Interests
‘Always happy to discuss these’
The Office, Dark, Chuck
Equities, ETFs, #Essentialism
Pihole, DNS resolvers, FOSS
Contact
- 1 DNA Way, South San Francisco, California 94080



